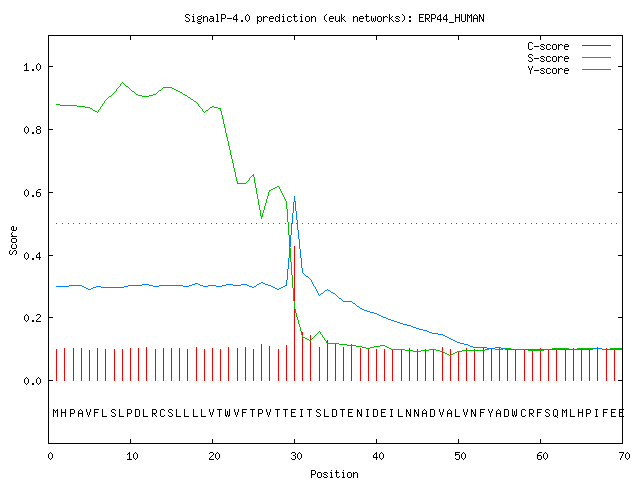
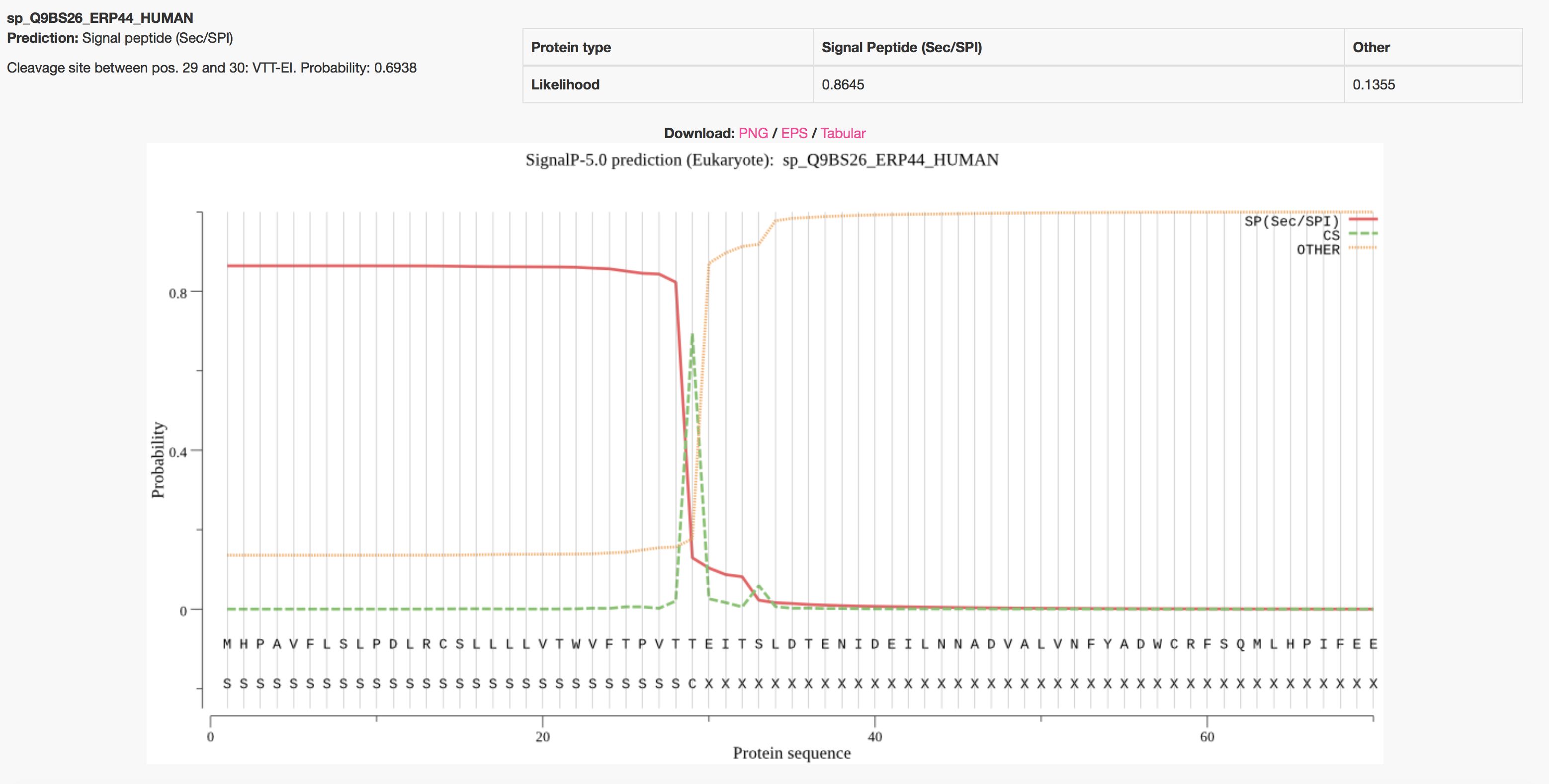
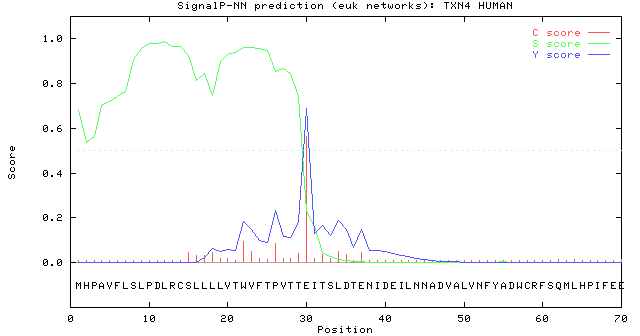
SignalP-5.0 prediction. Signal peptide likelihood was around 0.9505.... | Download Scientific Diagram

Sm29 signal peptide prediction using the SignalP 3.0 server according... | Download Scientific Diagram

SignalP 6.0 predicts all five types of signal peptides using protein language models | Nature Biotechnology

Proteomics reveals signal peptide features determining the client specificity in human TRAP-dependent ER protein import | Nature Communications

Proteome-Scale Screening to Identify High-Expression Signal Peptides with Minimal N-Terminus Biases via Yeast Display | ACS Synthetic Biology

USPNet: unbiased organism-agnostic signal peptide predictor with deep protein language model | bioRxiv

Signal Peptide Efficiency: From High-Throughput Data to Prediction and Explanation | ACS Synthetic Biology

SignalP-5.0 prediction. Signal peptide likelihood was around 0.9505.... | Download Scientific Diagram